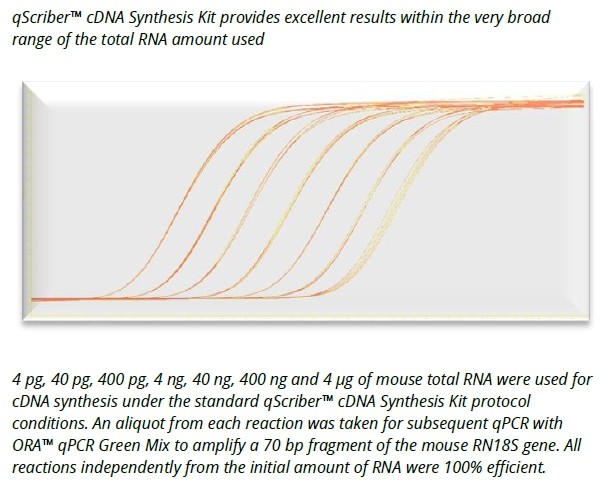
qScriber™ cDNA Synthesis Kit: Fast and reliable cDNA synthesis for gene expression analysis.
Applications
- cDNA template generation for qPCR or PCR.
- Unbiased, efficient cDNA synthesis.
- Detection of low target amounts.
- cDNA synthesis from complex templates.
Benefits
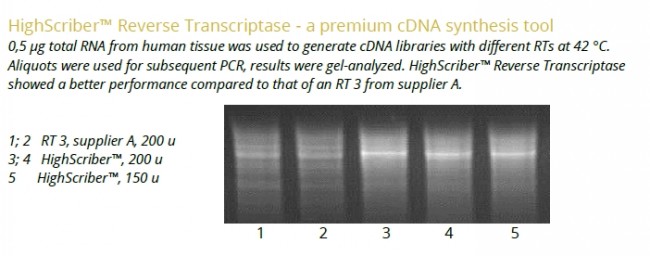
- Thermostable HighScriber™ Reverse Transcriptase blended with Ribonuclease Inhibitor for efficient cDNA synthesis.
- Optimized reaction mix with oligo (dT) and random primers for unbiased representation of mRNA ends.
- cDNA synthesis from complex templates at up to 55°C.
- High sensitivity detection from 1 pg of total RNA template.
The qScriber™ cDNA Synthesis Kit is a highly efficient and simple-to-use system for cDNA synthesis eliminating the need for tedious reaction optimization. The qScriber™ Enzyme Blend ensures high sensitivity detection from low copy number targets. The highly active and thermostable HighScriber™ Reverse Transcriptase blended with RNase Inhibitor allows for an efficient cDNA synthesis and reaction safety. The wide reaction temperature range (38°C - 55°C) ensures efficient transcription from GC rich templates.
cDNA (complementary or copy-DNA) synthesis is a fundamental technique used in both molecular biology research and in diagnostic applications. Reverse transcription process employs reverse transcriptase enzymes (RT), RNase inhibitors, template, random hexamer, oligo dT or specific primers and dNTPs. cDNA is produced aiming to perform different downstream applications. For example, for cloning purposes one takes specific primers, and performs the PCR after the cDNA synthesis to generate long DNA fragments for cloning into the plasmid vectors and for sequencing. For gene expression analysis purposes, short random primers are used, and generated short cDNA fragments unbiased represent the whole genome for qPCR analysis.
A good cDNA synthesis kit provides high yields of high-quality cDNA with minimal bias or variability. For complex targets containing high-GC regions thermostable reverse transcriptases are preferred, as they allow performing the reaction at higher temperature what prevents formation of secondary structures and increases the cDNA yield. Typically, the optimal cDNA synthesis temperature ranges between 37-42°C, depending on the reverse transcriptase enzyme used. Advanced RT enzymes go up to 55°C and above.
Optimizing cDNA synthesis protocol is crucial for generating accurate cDNA libraries. Several factors can influence the quality and yield of cDNA, including the amount and quality of RNA template, the concentration and quality of reverse transcriptase enzyme, quality of the primers, and the cDNA synthesis temperature.
At highQu, we offer several kits and enzymes for cDNA synthesis applications, these include: 1Step RT PCR Kit for fast PCR, HighScriber™ Reverse Transcriptase Mix for cloning purposes, and qScriber™ cDNA Synthesis Kit for a real-time qPCR-based gene expression analysis.
- Download Protocol and Specifications - Product Insert qScriber™ cDNA Synthesis Kit
- Ask for a Sample today
- Have technical questions? Contact us
- Download a list of Publications mentioning highQu products
- Or see how others use our products at bioz.com or google scholar
- Download Protocol and Specifications - Product Insert qScriber™ cDNA Synthesis Kit
- Download MSDS qScriber™ cDNA Synthesis Kit
- Need a lot-specific Certificate of Analysis? E-mail us at info@highQu.com
- Want custom formulations or bulk sizes? E-mail us at info@highQu.com, and check our OEM offers
- Have more specific questions? Contact us
- Download Protocol and Specifications - Product Insert qScriber™ cDNA Synthesis Kit
- Download MSDS qScriber™ cDNA Synthesis Kit
- Download highQu Catalogue of Premium Research Tools
- Download a list of Publications mentioning highQu products
- Or see how others use our products at bioz.com or google scholar
- Download Product Pricelist (DE 2021)
- Download all highQu Product Inserts and MSDS sheets
- Have questions? Contact us