Synthetic Carrier RNA (1 mg/ml or 10 mg/ml solution)
Applications
- All molecular biology applications where concentration of RNA or DNA solutions is required, such as:
- RNA extraction/isolation procedures
- DNA extraction/isolation procedures
- Clean-up and reprecipitation of RNA or DNA
Benefits
|
highQu Synthetic Carrier RNA is designed to be used in all kind nucleic acid purification and precipitation procedures as a carrier and co-precipitant of nucleic acids. It is especially useful to increase the amount of RNA or DNA pellet in low concentrated solutions, in such procedures, as viral RNA extraction from human specimen samples.
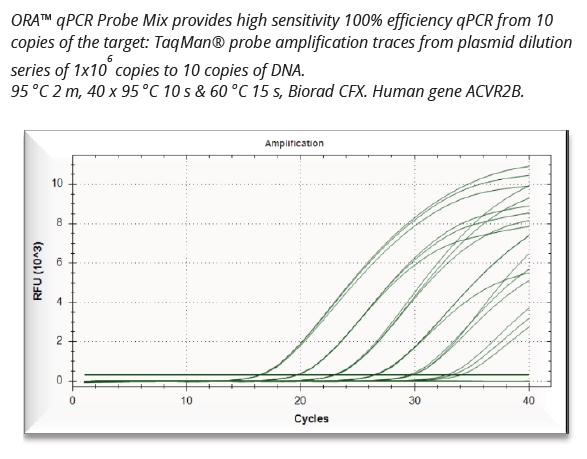
In contrast to commonly used carrier RNAs such as tRNA, yeast RNA, or sonicated salmon sperm DNA, the Synthetic Carrier RNA is free from animal or yeast RNA contamination. Coprecipitated RNA and DNA can be directly used for all common downstream applications, such as PCR or RT-PCR, as well as highly sensitive qPCR.
The use of carrier RNAs for coprecipitation of nucleic acids may interfere with spectrophotometrical concentration measurements.
The presence of carrier RNAs in the RNA or DNA solution may have some influence on certain enzymatic reactions performed by such enzymes that act on all nucleic acid molecules, for example T4 Polynucleotide Kinase or Terminal DNA Transferase.
Useful tips how to use coprecipitates such as carrier RNA for purification or concentration of PCR products and other low abundance DNA molecules without the use of PCR purification kits:
- Use an appropriate coprecipitant concentration: Too little coprecipitant may result in poor recovery of PCR fragment, while too much can lead to increased background noise when performing concentration measurements.
- Use Synthetic Carrier RNA in DNA, PCR reaction product or RNA solutions during alcohol precipitation step.
- To maximize the yield of nucleic acids, before adding salt (Sodium acetate) and Ethanol or Isopropanol, first add Synthetic Carrier RNA and mix it well with DNA/RNA sample or PCR product.
Use following amounts of Synthetic carrier RNA:
- Recommended final concentration in precipitation solution is 10–20 μg/ml
- For example, add 1 μl of 10 mg/ml Synthetic Carrier RNA into 200 μl of RNA or DNA sample which will be precipitated with 3 x volumes of ethanol.
- Alternatively, add 5 μl of 1 mg/ml Synthetic Carrier RNA into 100 μl of RNA or DNA sample which will be precipitated with 3 x volumes of ethanol.
- Incubate at low temperatures: After adding the coprecipitant and precipitation reagents, incubate the reaction mixture at a low temperature, usually -20°C or lower, for an adequate duration (e.g., 30 minutes to overnight).
For best purification results, and highest purity PCR purification product, wash the precipitated DNA pellet: After centrifugation and removal of the supernatant, wash the DNA pellet with ice-cold 70% ethanol to remove residual impurities and dry the pellet before resolving it in nuclease free water.
- Download Protocol and Specifications - Product Insert Synthetic Carrier RNA
- Ask for a Sample today
- Have technical questions? Contact us
- Download a list of Publications mentioning highQu products
- Or see how others use our products at bioz.com or google scholarhttps://www.highqu.com/media/wysiwyg/ressources/manuals/SCR01_Synthetic_Carrier_RNA_PI.pdf
- Download Protocol and Specifications - Product Insert Synthetic Carrier RNA
- Download MSDS Synthetic Carrier RNA
- Need a lot-specific Certificate of Analysis? E-mail us at info@highQu.com
- Want custom formulations or bulk sizes? E-mail us at info@highQu.com, and check our OEM offers
- Have more specific questions? Contact us
- Download Protocol and Specifications - Product Insert Synthetic Carrier RNA
- Download MSDS Synthetic Carrier RNA
- Download highQu Catalogue of Premium Research Tools
- Download a list of Publications mentioning highQu products
- Or see how others use our products at bioz.com or google scholar
- Download Product Pricelist (DE)
- Download all highQu Product Inserts and MSDS sheets
- Have questions? Contact us